Labelbox•September 3, 2021
New study finds that non-expert labeling teams can create high quality training data for medical use cases
Producing high quality training data for any ML use case can be costly, and it’s doubly so for specialized use cases that require experienced domain experts to review (and often label) training data. In the agriculture industry, for example, ML teams training models to recognize weeds or diseased crops need to have their training data examined and corrected by trained agronomists and horticulturists, whose time is not only expensive, but also limited.
Those developing algorithms for medical use cases face the same challenge. Having doctors meticulously label training data, or even review labeled data, can be an arduous process. If there was a way to create high quality training data with less involvement from physicians, ML teams could produce life saving algorithms at a fraction of the cost — and a faster rate.
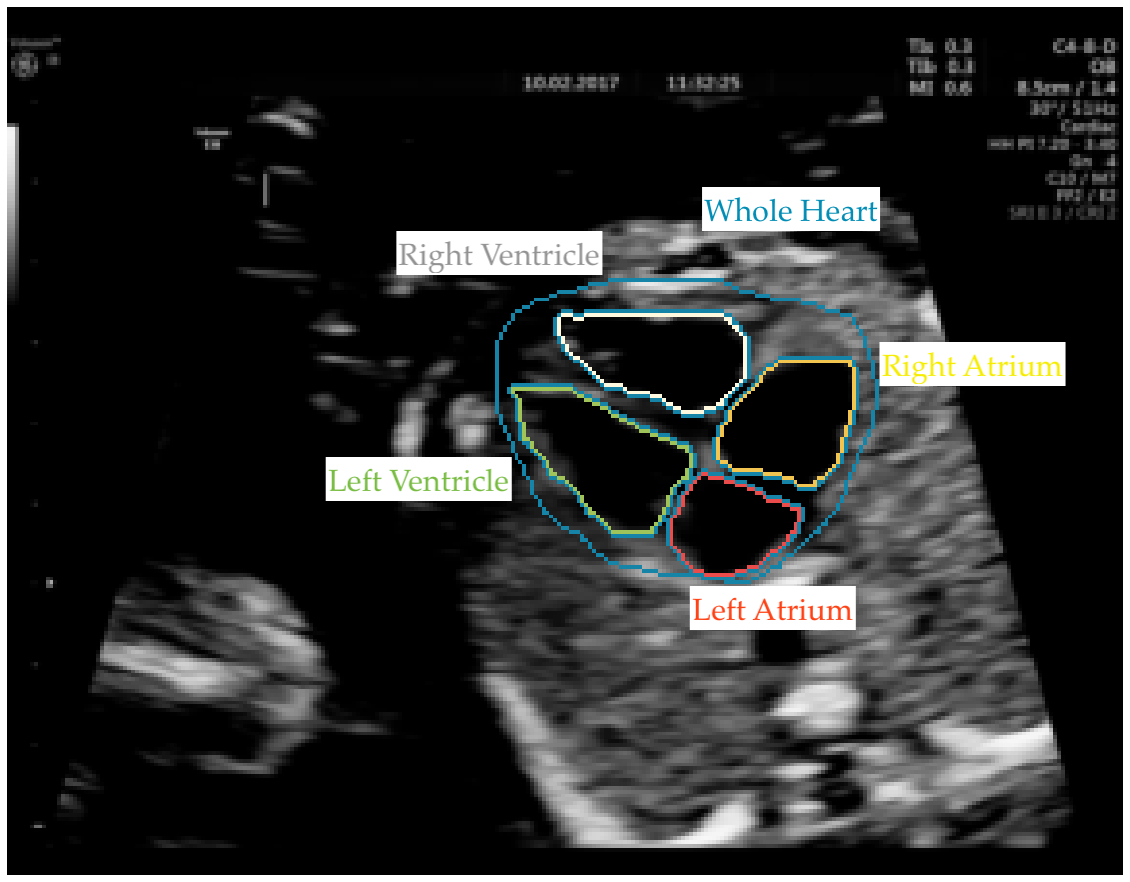
A new study spearheaded by researchers from Imperial College London’s BioMedIA research group and the iFind Project set out to challenge the widely held assumption that only those with years of training in medicine can provide quality annotations for use in clinical deep learning models. The team partnered with Labelbox and our Labeling Operations team to conduct the experiment, where minimally trained, novice labeling workforces were tasked with annotating fetal ultrasound imagery for signs of a specific congenital heart disease: hypoplastic left heart syndrome.
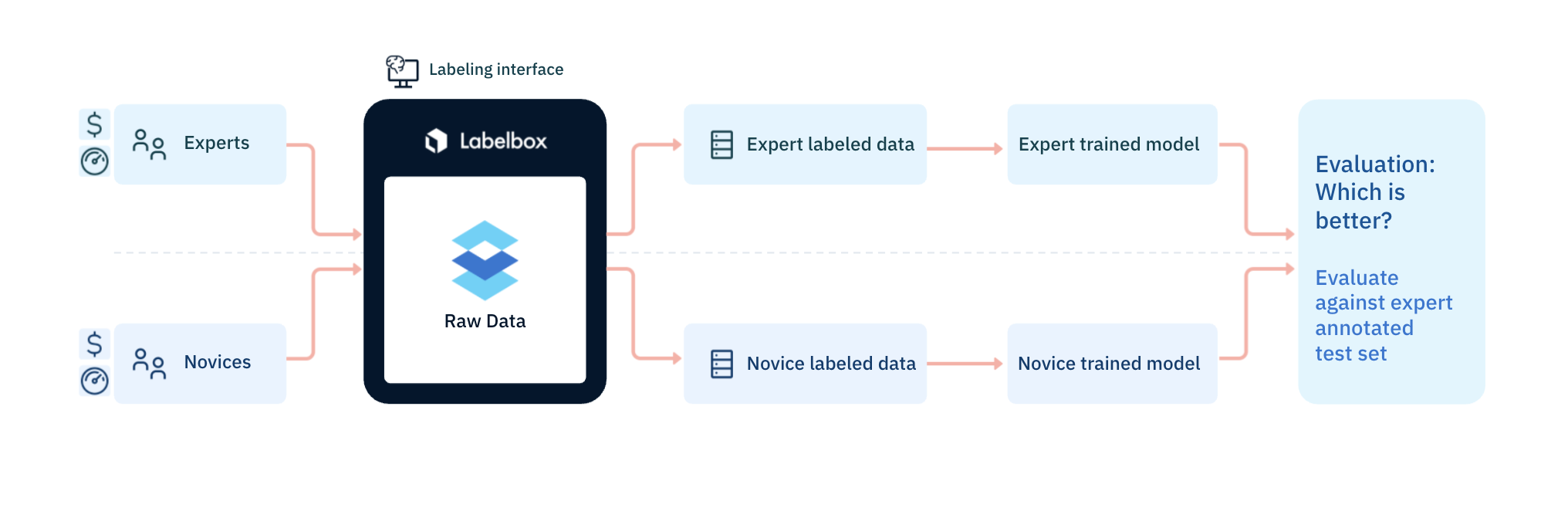
Researchers then trained the same algorithm on two datasets: one labeled by the Labelbox workforces and one labeled by experts in medical imagery. Throughout the experiment, the team tracked benchmark and consensus statistics to ensure the quality of the labels created before performing their own, more detailed analyses. Because they partnered with Labelbox, they were able to set up their experiment and datasets quickly and easily, as well as choose from a range of annotation types.
“Our experts found these straightforward to use and were able to annotate data very quickly to a high standard,” said Imperial College London researcher Sam Budd.
The experiment found that these novice annotators could perform complex medical image segmentation tasks to a high standard. Their labels were similar in quality to those provided by experts, and prediction performance in the models were comparable in both image segmentation and downstream classification tasks.
"We've challenged the assumption that only experts can do medical image annotation and found that novices at Labelbox can do well with just a few pages of instructions and examples," said Budd in a tweet announcing the study's results.
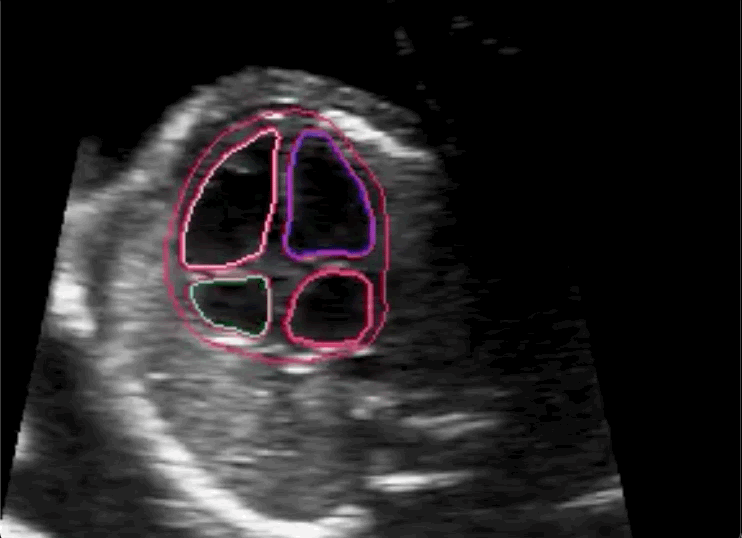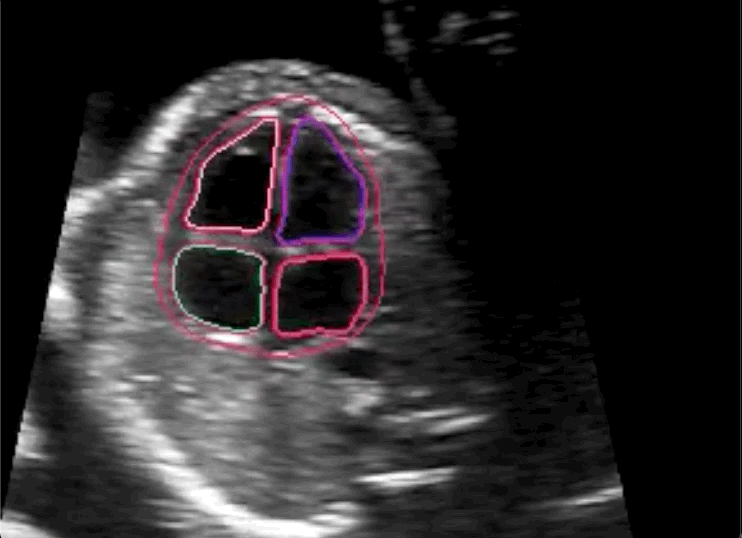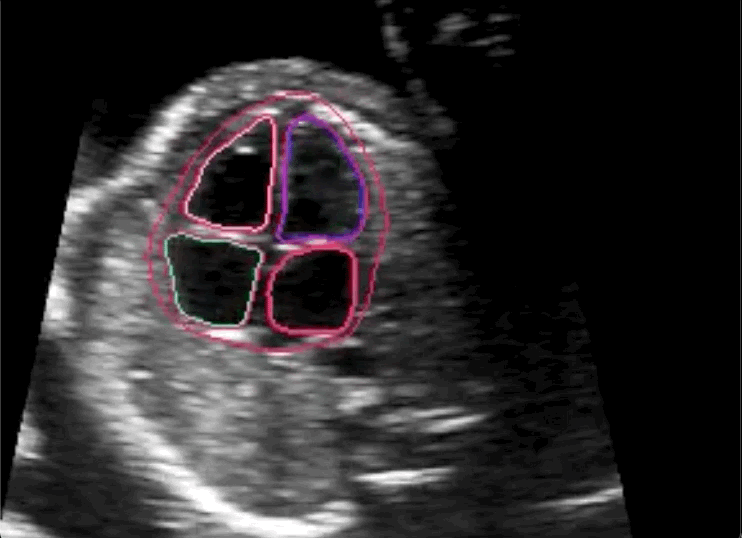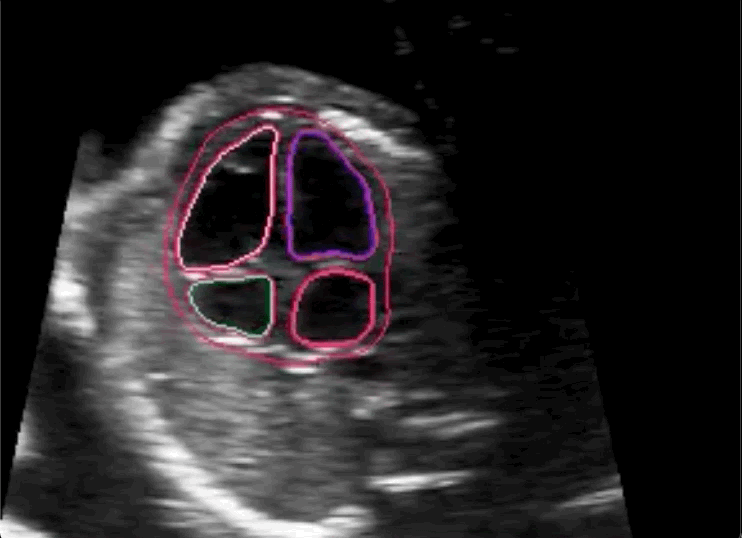
The researchers concluded in their paper that novice labeling teams can play a vital role in developing high performing models in specialized medical use cases when employed in combination with existing methods to handle noisy annotations and choose the most informative annotations for the model.
“The speed of the Labelbox Workforce team was impressive and everything was annotated within our expected time-frame, allowing us to parse the results and perform the necessary experiments in time to submit a paper that was accepted into the FAIR workshop at MICCAI 21. I would recommend Labelbox to peers and others in the research community who have data that needs labelling but not the manpower to do so themselves — by providing some simple instructions we were able to acquire sufficient quality labels to train high-performing computer vision models for our purposes.”
— Sam Budd, PhD Student & Researcher at BioMedIA at Imperial College London
You can read the research paper on the study here. To learn more about how you can work with the Labelbox Workforce to annotate your training data accurately and efficiently, visit our labeling services page.

 All blog posts
All blog posts

