Automated detection of malaria parasites using convolutional neural networks
Summary: Researchers from Imperial College London recently focused on developing an automated image analysis method to improve accuracy and standardization of smear inspection that retains capacity for expert confirmation and image archiving.
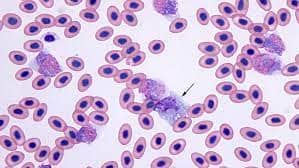
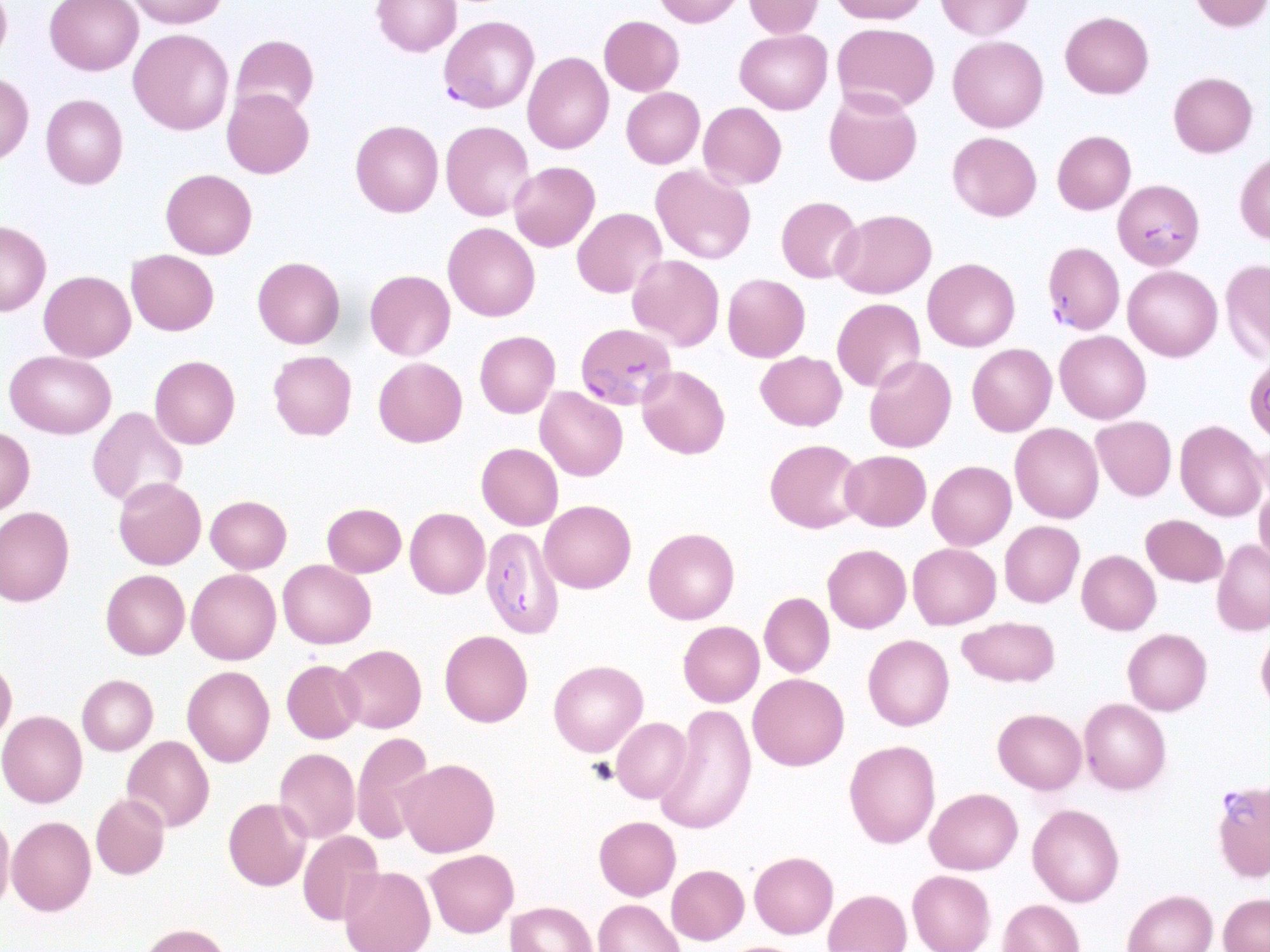
Challenge: Microscopic examination of blood smears is known to be the gold standard for laboratory inspection and diagnosis of malaria. Smear inspection is, however, time-consuming and dependent on trained microscopists with results varying in accuracy. To advance this technique using convolution neural networks (CNNs), a machine learning method was developed to hone in on red blood cell (RBC) detection, differentiation between infected/uninfected cells, and parasite life stage categorization from unprocessed, heterogeneous smear images.
Findings: Based on a pretrained Faster Region-Based Convolutional Neural Networks (R-CNN) model for RBC detection, their model performed accurately, with an average precision of 0.99 at an intersection-over-union threshold of 0.5. Application of a residual neural network-50 model to infected cells also performed accurately, with an area under the receiver operating characteristic curve of 0.98.
Combined with a mobile-friendly web-based interface that they built, called PlasmoCount, their ML method permits rapid navigation through and review of results for quality assurance. By standardizing assessment of Giemsa smears, their method markedly improves inspection reproducibility and presents a realistic route to both routine lab and future field-based automated malaria diagnosis.
How Labelbox was used: The researchers leveraged a model-assisted approach for labeling using the Labelbox platform which greatly improved labeling speed and performance.
Each labeling round contained ~100 raw images of Giemsa smears. To aid the first labeling round, they trained their object detection model on a dataset of Plasmodium vivax-infected blood smears from the Broad Bioimage Benchmark Collection. Predictions on their P. falciparum dataset were then uploaded as prelabels using the Labelbox Python SDK. For each of the following labeling rounds, the RBC detection model and malaria identification classifier were trained on all the previous labeled datasets to generate new labels.
Annotators could then correct, add, and delete bounding boxes around each RBC, and choose from three labels: infected, uninfected, and unsure. All images were labeled five times by five designated annotators from three different research centers for the test set.
You can read the full PDF here.

 All posts
All posts